Of Viruses and Lego Bricks
The Lego Death Star and the Brick Testament use the same building blocks. Viruses are like that too
Although my primary research revolves around the beautiful capsid proteins that protect polyomavirus genomes from the outside world, I’ve also developed a bit of a side-hustle hunting exotic viruses. This has typically involved directly sequencing samples collected in places like supermarkets. When the pandemic forced me to pursue remote work, I began eyeballing the Sequence Read Archive (SRA) - a massive public repository where biologists deposit their raw sequencing data. It took me four long years to make sense of the evolutionary chaos I witnessed down in the SRA datamines, but I’m proud to report that the manuscript is finally finished and under review in the exciting new vetoless track at eLife.
In this initial post expanding upon the datamining results, I’ll focus on virus taxonomy. Taxonomy might sound like a pretty esoteric place to start, but my feeling is that Ursula LeGuin was wrestling with an essential truth in her Earthsea novels:
My name, and yours, and the true name of the sun, or a spring of water, or an unborn child, all are syllables of the great word that is very slowly spoken by the shining of the stars. There is no other power…. For magic consists in this, the true naming of a thing
Humankind can have little hope of mastering a thing if we don’t at least have a succinct name to call it by.
In a seminal 2015 analysis, Koonin and colleagues described viruses as “the ultimate modularity.” In contrast to animals, which inherit genes vertically from our ancestors, viruses often appropriate genes horizontally - stealing either from the host cell or from completely unrelated viruses. The emerging model is that viruses are sort of like mix-and-match bags of Lego bricks, particularly over million-year time scales.
Although traditional Linnaean taxonomy works well for classifying organisms that evolve through vertical inheritance, horizontal modularity basically plants a bomb in Linnaeus’s system. To illustrate the problem, let’s imagine a world in which mythological sphinxes actually exist. Sphinxes can’t be meaningfully classified either as simply a type of human or simply a type of lion - and designating them as a third, entirely separate taxon would fail to acknowledge important similarities to the other two groups.
The real-world existence of sphinxes would force us to develop separate taxonomies for heads and bodies - enabling us to think of a primate-carnivore chimera as a “primarnivore.” A real-world virological application of the principle is the awkwardly named Japanese eel endothelial cells-infecting virus (JEECV) - a lethal pathogen that’s currently pushing endangered eels toward extinction. The capsid proteins of JEECV resemble the virion proteins of adenoviruses, while its DNA-replication proteins resemble those of polyomaviruses. The field has settled on the family name “adomavirus” for this class of viruses.
For a more playful thought experiment, let’s imagine traveling back in time to show Aristotle a mountain bike. Aristotle would undoubtedly recognize the similarities between bikes and familiar two-wheeled chariots, and he would likely be able to quickly infer the functions of the mountain bike’s unfamiliar handlebars and shock absorbers. If we next present Aristotle with a pogo stick, he would quickly recognize it as a shock absorber with handlebars. Although pogo sticks were invented a century after bikes, it’s incorrect to imagine that pogo sticks descended from chariots in a stepwise process involving a mountain bike intermediate. Vehicles, like viruses, are more accurately understood as modular amalgamations of discrete inventions that can be recombined horizontally.
Now let’s imagine showing Aristotle a helicopter. He might initially guess that the blades on top could be some sort of weapon - unless he were told the vehicle is called a “helicopter” and the blade array is called a “propeller.” It wouldn’t be helpful to tell Aristotle that the device is named “unclassified vehicle” and the blade array is named “hypothetical part.” Names are important!
For modular organisms, it’s especially important to have clear names for hallmark gene categories. Unfortunately, the virology community is shockingly bad at this task. For example, an important adenovirus oncogene I’ve settled on calling “Oncorf6” is represented in the literature by various combinations of the terms hypothetical protein, early E4, protein 6, control protein, 15.9 kDa, 16 kDa, 19.8 kDa, 32 kDa, 33.2 kDa, 34 kDa, 34.1 kDa, 34.6 kDa, 34.7 kDa, 34K, 34K-2, E4.2, E4-1, E4-3, E4-6, BAdVBgp27, ORFD, ORFE, ORF3, ORF5, ORF6, ORF6/7, ORF26, 245R, 253R, or E4orf6.
It would be very useful if the International Committee on Taxonomy of Viruses or committees organized by major sequence repositories could suggest standard names for key viral genes. The good news on this front is that AI tools like AlphaFold and RoseTTAFold have suddenly made it much easier to figure out which genes encode structurally similar proteins. For example, a gene name I coined while trying to make sense of the datamines is “Cah,” which stands for predicted capsid-surface protein with alpha-helix-rich predicted fold.
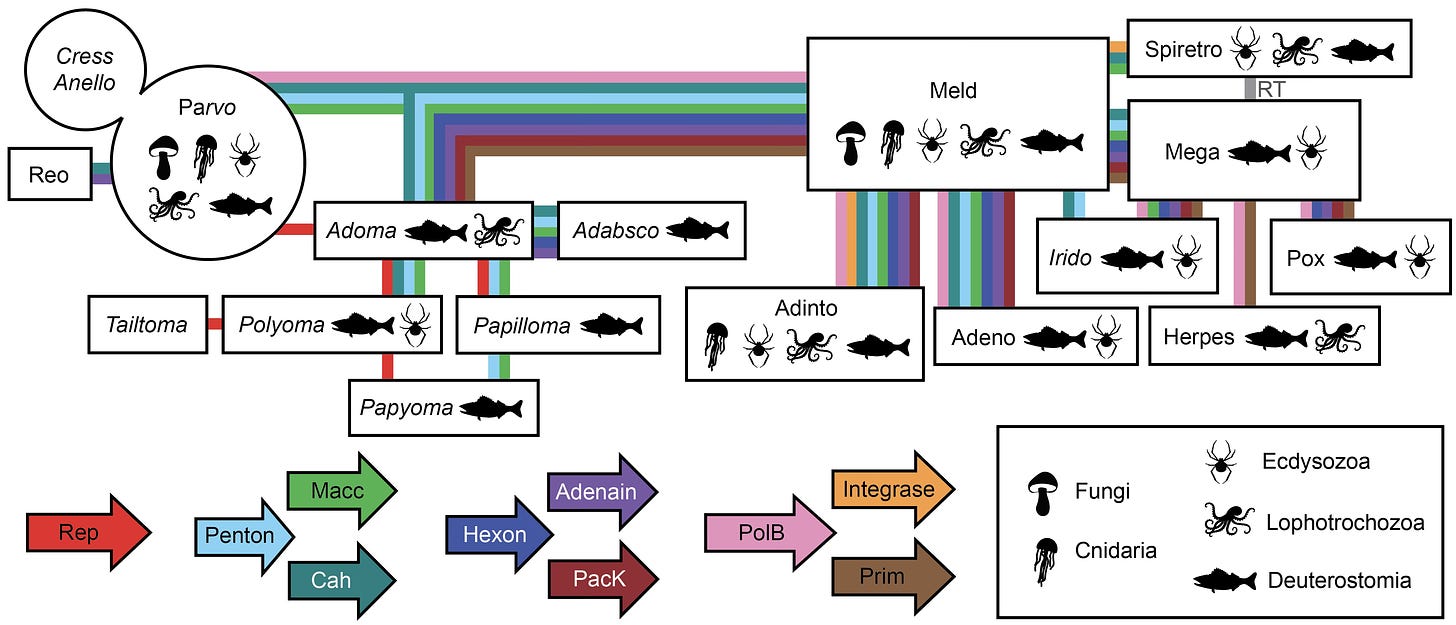
Another bedrock taxonomic problem is that it’s hard to represent the evolution of modular organisms using traditional phylogenetic trees. The datamining project made extensive use of all-against-all network analyses, which I’m a big fan of. Subway map representations are another good way to portray the horizontal flow of different gene classes between virus groups. Here’s my attempt to summarize the inferred flow of key genes among the thousands of exotic viruses I saw in the datamines.
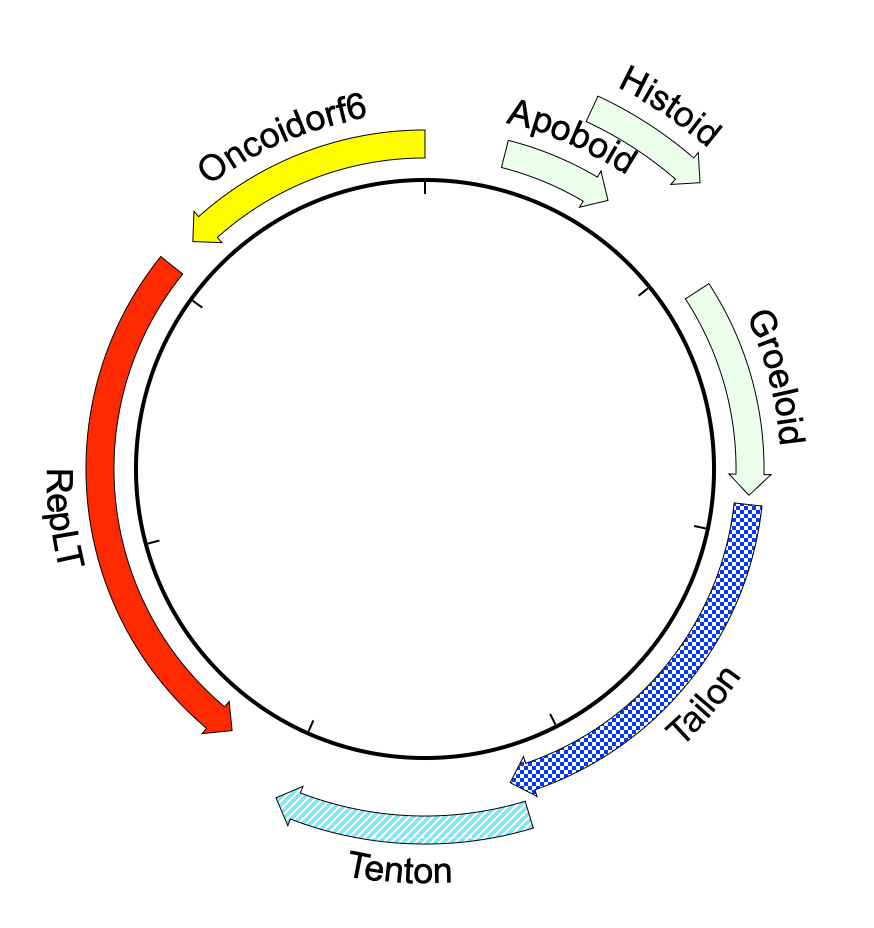
At a practical level, the most important aspect of the gene-centric view of virus evolution is that it opens up a powerful new hunting method. A favorite example, found in seawater off the coast of Tierra del Fuego, has an easily recognizable homolog of the familiar polyomavirus Large Tumor antigen (RepLT) horizontally recombined with capsid proteins from outer space. Even with AlphaFold, it’s hard to understand the evolutionary origins of the Tailon and Tenton proteins. The nearest structural similarities are found in viruses that infect archaea, but the signal is very faint. In a world with sphinxes, you can discover unfamiliar lion bodies simply by listening for talking heads!
For better or worse, the world only has one of me - and now that the pandemic is becoming more manageable I’ve been reverting back to my day job as a vaccine developer. My next post will cover my dreams for how somebody might be able to democratize these new virus-hunting approaches so others can trawl the depths and find even stranger sphinxes than I did. Then I can make vaccines against them. Just let me know.




Can we be certain the seemingly chimeric sequences are due to horizontal transfer and not convergent evolution?